PiBlue™ Phosphate Assay Kit
Application
- For sensitive and high-throughput phosphate determination.
Key Features
- Reagent very stable. Due to our innovative formulation, no precipitation of reagent occurs. Therefore no filtration of reagent is needed prior to assays, as is often required with other commercial kits.
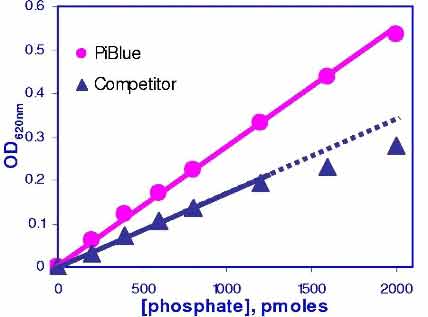
- High sensitivity and wide detection range: detection of as little of 20 pmoles of phosphate and useful range between 0.4 µM and 50 µM phosphate.
- Fast and convenient: single reagent “mix-and-measure” assay allows quantitation of free phosphate within 30 minutes.
- Compatible with routine laboratory and HTS formats: assays can be performed in tubes, cuvettes or microplates, on spectrophotometers and plate readers.Robust and amenable to HTS: Z factors of 0.7 to 0.9 are observed in 96-well plates. Can be readily automated on HTS liquid handling systems
Method
- OD620nm (malachite green)
Samples
- Enzyme reactions with free phosphate releases
Species
- All
Procedure
- 30 min
Size
- 500 tests
Detection Limit
- 0.4 µM
Shelf Life
- 12 months
More Details
The PiBlue™ Phosphate Assay Kit is based on a proprietary formulation of the malachite green dye. The PiBlue™ reagent forms a blue colored complex with free orthophosphate. The rapid color formation from the reaction can be conveniently measured on a spectrophotometer (600 – 660 nm) or on a plate reader. The non-radioactive colorimetric assay kits have been optimized to offer superior sensitivity and prolonged shelf life. The assay is simple and fast, involving a single addition step for phosphate determination. Assays can be performed in tubes, cuvettes or multi-well plates. The assays can be conveniently executed in 96-well plates for high-throughput screening of enzyme inhibitors.What is the difference between your 3 phosphate assays: DIPI-500, POMG-25H, POPB-500?
All three products are based on the malachite green dye. While the PiBlue (POPB-500) and QuantiChrom (DIPI-500) Phosphate Assay Reagents are identical, they differ in the standards. The PiBlue is designed for customers who want prepare a full calibration curve for their assays, and the latter has one set of a blank control and 30 µM phosphate standard. The reagent is more diluted than the reagent used in the Malachite Green Phosphate Assay Kit (POMG-25H). As a result, one adds 100 µL reagent (POPB-500 or DIPI-500) to 50 µL sample, whereas for the Malachite Green based assay, one adds 20 µL Reagent to 80 µL sample.
In practice, the DIPI-500 is the most convenient kit to use, because there is no need to prepare a calibration curve there is no need for extra mixing. Because adding a larger volume (100 μL) to a small volume (50 μL) effects efficient mixing in well, no extra mixing is needed. The POPB-500 requires preparation of calibration curve. With the POMG-25H Kit, one needs to mix the reagent with the sample, because adding 20 µL to 80 µL does not mix efficiently by itself. Also a preparation of a standard curve is necessary.
I’m using the Malachite Green Phosphate Assay Kit (POPB-500) to detect GTPase activity. However, I saw high background and no increase in phosphate liberation.
This assay is very sensitive to free phosphate. Before each assay, it is important to check that all materials (substrates, enzyme preparations and buffers) are pure and free of contaminating phosphate. This can be conveniently done by adding 20 µL of the Working Reagent to 80 µL sample solution. The blank OD values at 620 nm should be lower than 0.2. If the OD readings are higher than 0.2, check water phosphate level. Double distilled water usually have OD readings lower than 0.1. Lab detergents may contain high levels of phosphate. Make sure that lab wares are free from contaminating phosphate after thorough washes.
In the case of enzyme reactions, the reaction may be terminated directly by the addition of the acidic Working Reagent. Dilution of reaction mixture may be necessary prior to the assay. For ATPase or GTPase assays, the ATP or GTP concentration should be lower than 0.25 mM. If the reaction mixture contains > 0.25mM ATP or GTP, dilute samples in distilled water. For example, if the ATPase reaction contained 1 mM ATP, at the end of reaction dilute reaction mixture 4-fold in water prior to the assay.
The chromogenic reaction is completed within 30 min at room temperature. Read OD at or shorter than 30 min to minimize background due to slow hydrolysis of the substrates (ATP or GTP) by the acidic reagent.
If the background OD is in the normal range, but there is still no free phosphate detected, it is likely the GTPase was not active. To confirm this possibility, one could run a test without the enzyme by spiking free phosphate (e.g. 0.5 – 40 µM) into the mixture of Buffer and Substrate. In this case, one should see an increase in OD with increasing free phosphate concentrations. This confirms that the enzyme was not active in the assay.
Christensen, T., et al. (2020). In vitro characterization of a novel consensus bacterial 6-phytase and one of its variants. Current Biochemical Engineering 6(3): 156-171. Assay: Phosphate in Trichoderma reesei phytase.
Nitta, K., et al. (2021). Investigation of the effects of actinorhodin biosynthetic gene cluster expression and a rpoB point mutation on the metabolome of Streptomyces coelicolor M1146. Journal of Bioscience and Bioengineering. Assay: Phosphate in Streptomyces coelicolor M1146 cells.
Nitta, K., et al. (2020). Multi-omics analysis of the effect of cAMP on actinorhodin production in Streptomyces coelicolor. Frontiers in Bioengineering and Biotechnology 8: 595552. Assay: Phosphate in bacteria nutrition medium.
Lejeune, C., et al. (2021). Impact of phosphate availability on membrane lipid content of the model strains, Streptomyces lividans and Streptomyces coelicolor. Frontiers in Microbiology 12: 623919. Assay: Phosphate in R2YE agar medium.
Hwang, J. D., Ortiz-Maldonado, M., & Paramonov, S. (2016). Delivery of Formulated Industrial Enzymes with Acoustic Technology. Journal of laboratory automation, 21(1), 153-165. Assay: Phosphate in Buttiauxella sp Phytase.
Demidenko AA et al (2011). Effects of viscogens on RNA transcription inside reovirus particles. J Biol Chem. 286(34):29521-30. Assay: Phosphate in rat lipid.
Liu Q et al (2011). Competition between foliar Neotyphodium lolii endophytes and mycorrhizal Glomus spp. fungi in Lolium perenne depends on resource supply and host carbohydrate content. Functional Ecology 25:910-920. Assay: Phosphate in mouse intracellular phosphate.
Lopez-Vales R, et al (2010). Fenretinide promotes functional recovery and tissue protection after spinal cord contusion injury in mice. J Neurosci. 30(9):3220-6. Assay: Phosphate in mouse lipid.
Oborna I et al (2010). Increased lipid peroxidation and abnormal fatty acid profiles in seminal and blood plasma of normozoospermic males from infertile couples. Hum Reprod. 25(2):308-16. Assay: Phosphate in fish marine teleost follicles, eggs.
Oborna I et al. (2010). Increased lipid peroxidation and abnormal fatty acid profiles in seminal and blood plasma of normozoospermic males from infertile couples. Hum Reprod. 25(2):308-16. Assay: Phosphate in human blood, semen.
Polewski MD, et al (2010). Inorganic pyrophosphatase induces type I collagen in osteoblasts. Bone 46(1):81-90. Assay: Phosphate in mouse/human cellular extract.
Cramp RL et al (2009). Ups and downs of intestinal function with prolonged fasting during aestivation in the burrowing frog, Cyclorana alboguttata. J Exp Biol. 212(22):3656-63. Assay: Phosphate in mouse lipid.
Cramp RL, et al (2009). Ups and downs of intestinal function with prolonged fasting during aestivation in the burrowing frog, Cyclorana alboguttata. J Exp Biol. 212(22):3656-63. Assay: Phosphate in frog Na+/K+-ATPase extract.
Huxtable AG, et al (2009). Tripartite purinergic modulation of central respiratory networks during perinatal development: the influence of ATP, ectonucleotidases, and ATP metabolites. JNeurosci. 29(47):14713-25. Assay: Phosphate in rat ATPase extract.
Kang HY et al (2008). Altered TNSALP expression and phosphate regulation contribute to reduced mineralization in mice lacking androgen receptor. Mol Cell Biol.28(24):7354-67. Assay: Phosphate in rat ATPase extract.
Kang HY, et al (2008). Altered TNSALP expression and phosphate regulation contribute to reduced mineralization in mice lacking androgen receptor. Mol Cell Biol.28(24):7354-67. Assay: Phosphate in mouse cytosolic fraction.
Chen X et al (2007). The isolation and structure of membrane lipid rafts from rat brain. Biochimie 89(2):192-6. Assay: Phosphate in mouse/human cellular extract.
Chen X, et al (2007). The isolation and structure of membrane lipid rafts from rat brain. Biochimie 89(2):192-6. Assay: Phosphate in rat lipid.
Hough TA et al (2007). Novel mouse model of autosomal semidominant adult ypophosphatasia has a splice site mutation in the tissue nonspecific alkaline phosphatase gene Akp2. J Bone Miner Res. 22(9):1397-407. Assay: Phosphate in frog Na+/K+-ATPase extract.
Hough TA, et al (2007). Novel mouse model of autosomal semidominant adult hypophosphatasia has a splice site mutation in the tissue nonspecific alkaline phosphatase gene Akp2. J Bone Miner Res. 22(9):1397-407. Assay: Phosphate in mouse intracellular phosphate.
Lau K.H. et al. (2006). An osteoclastic protein-tyrosine phosphatase is a potential positive regulator of the c-Src protein-tyrosine kinase activity: a mediator of osteoclast activity. J Cell Biochem. 97(5):940-55. Assay: Phosphate in rabbit phosphatases.
To find more recent publications, please click here.
If you or your labs do not have the equipment or scientists necessary to run this assay, BioAssay Systems can perform the service for you.
– Fast turnaround
– Quality data
– Low cost
Please email or call 1-510-782-9988 x 2 to discuss your projects.
$279.00
For bulk quote or custom reagents, please email or call 1-510-782-9988 x 1.
Orders are shipped the same day if placed by 2pm PST
Shipping: RT
Carrier: Fedex
Delivery: 1-2 days (US), 3-6 days (Intl)
Storage: 4°C upon receipt
Related Products
You may also like…
| Name | SKU | Price | Buy |
|---|---|---|---|
| Malachite Green Phosphate Assay Kit | POMG-25H |
$329.00 |
Why BioAssay Systems
Quality and User-friendly • Expert Technical Support • Competitive Prices • Expansive Catalogue • Trusted Globally
