QuantiChrom™ Ethanol Assay Kit
Application
- For quantitative determination of ethanol and alcohol metabolism.
Key Features
- Sensitive and accurate. Detection range 0.04 – 4% alcohol in 96-well plate assay.
- Convenient and high-throughput. The procedure involves adding a single working reagent, incubation for 8 min, adding a Stop Reagent, and reading the optical density. Can be readily automated as a high-throughput 96-well plate assay for thousands of samples per day.
- Versatility. Assays can be executed in 96-well plates or cuvet.
Method
- OD580nm (Chemical)
Samples
- Alcoholic beverages etc
Species
- All
Procedure
- 10 min
Size
- 500 tests
Detection Limit
- 0.0004
Shelf Life
- 12 months
More Details
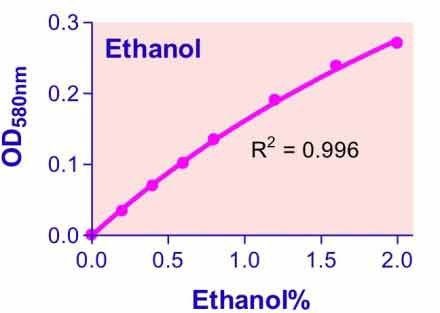
Alcoholic drinks are among the daily consumed beverages. Studies have shown heavy alcohol consumption may lead to various forms of liver diseases and to increased mortality rates. Quantitative determination of alcohol (ethanol, C2H5OH) finds applications in basic research, drug discovery, clinic studies, and winery. Simple, direct, and automation-ready procedures for measuring ethanol concentration are very desirable. BioAssay Systems QuantiChrom™ ethanol assay kit is based on an improved dichromate method, in which dichromate is reduced by ethanol to a bluish chromic (Cr3+) product. The intensity of color, measured at 580 nm, is a direct measure of the alcohol concentration in the sample. The optimized formulation substantially reduces interference by substances in the raw samples and exhibits high sensitivity.We are working with a very small murine blood sample (20 μl) and cannot safely remove more than this for our purposes. Will your assay work with such a small sample? Would this require any special methods?
Serum samples will be diluted 3-fold during deproteination. You have to add water to give 100 μL sample for the assay. The ethanol concentration should be 0.04% or higher in the diluted sample, or (>0.2%) in the initial serum sample, in order to be quantified accurately. Our enzyme based ethanol assay (ECET-100), which is more sensitive, might be a better option for you.
Does glucose interfere with the assay?
Yes, glucose and other compound containing hydroxyl groups (sugars, glycerol, etc.) do interfere with the assay. Samples containing high amounts of glucose should be pretreated with our saccaride removal kit (DSRK-500). Our enzyme based ethanol assay (ECET-100), which is more specific, is the better option for such samples.
Sawant, O. B., et al. (2020). Prenatal alcohol exposure and maternal glutamine supplementation alter the mTOR signaling pathway in ovine fetal cerebellum and skeletal muscle. Alcohol (Fayetteville, N.Y.), 89, 93-102. Assay: Ethanol in sheep blood.
Laksitorini, M. D., et al. (2020). Impact of Wnt/beta-catenin signaling on ethanol-induced changes in brain endothelial cell permeability. Journal of Neurochemistry. Assay: Ethanol in mouse tissue and human cells.
Nugent, B., Ali, S. S., Mullins, E., & Doohan, F. M. (2019). A Major Facilitator Superfamily Peptide Transporter From Fusarium oxysporum Influences Bioethanol Production From Lignocellulosic Material. Frontiers in microbiology 10:295. Assay: Ethanol in Fungus.
Contreras, M. L., de la Fuente-Ortega, E., Vargas-Roberts, S., Munoz, D. C., Goic, C. A., & Haeger, P. A. (2017). NADPH Oxidase Isoform 2 (NOX2) is involved in drug addiction vulnerability in progeny developmentally exposed to ethanol. Frontiers in Neuroscience, 11, 338. Assay: Ethanol in Sprague Dewley rats serum.
Duy, D. L., Suda, Y., & Irie, K. (2017). Cytoplasmic deadenylase Ccr4 is required for translational repression of LRG1 mRNA in the stationary phase. PloS one, 12(2), e0172476. Assay: Ethanol in E. coli media.
Kim HR, et al (2011). Raw starch fermentation to ethanol by an industrial distiller’s yeast strain of Saccharomyces cerevisiae expressing glucoamylase and alpha-amylase genes. Biotechnol Lett. 33(8):1643-8. Assay: Ethanol in yeast starch.
Pisu MG, et al (2011). Effects of voluntary ethanol consumption on emotional state and stress responsiveness in socially isolated rats. Eur Neuropsychopharmacol.21(5):414-25. Assay: Ethanol in rat plasma.
Alam MZ,et al (2009). Production of bioethanol by direct bioconversion of oil-palm industrial effluent in a stirred-tank bioreactor. J Ind Microbiol Biotechnol. 36(6):801-8. Assay: Ethanol in plant oil-palm.
Lin, YY et al (2009). Protein acetylation microarray reveals that NuA4 controls key metabolic target regulating gluconeogenesis. Cell 136(6):1073-84. Assay: Ethanol in yeast culture medium.
Wang J, et al (2009). Altered pattern of Na,K-ATPase activity and mRNA during chronic alcohol consumption by juvenile and adolescent rats. Cell Mol Neurobiol. 29(1):69-80. Assay: Ethanol in rat blood.
Biggio F, et al (2007). Flumazenil selectively prevents the increase in alpha(4)-subunit gene expression and an associated change in GABA(A) receptor function induced by ethanol withdrawal. J Neurochem. 102(3):657-66. Assay: Ethanol in rat primary neuronal cultures, medium.
Khanna D, et al (2007). Inducible nitric oxide synthase attenuates adrenergic signaling in alcohol fed rats. J Cardiovasc Pharmacol. 50(6):692-6. Assay: Ethanol in rat serum.
To find more recent publications, please click here.
If you or your labs do not have the equipment or scientists necessary to run this assay, BioAssay Systems can perform the service for you.
– Fast turnaround
– Quality data
– Low cost
Please email or call 1-510-782-9988 x 2 to discuss your projects.
$549.00
For bulk quote or custom reagents, please email or call 1-510-782-9988 x 1.
Orders are shipped the same day if placed by 2pm PST
Shipping: RT
Carrier: Fedex
Delivery: 1-2 days (US), 3-6 days (Intl)
Storage: 4°C upon receipt
Related Products
You may also like…
| Name | SKU | Price | Buy |
|---|---|---|---|
| QuantiChrom™ Alcohol Dehydrogenase Assay Kit | DADH-100 |
$459.00 |
Why BioAssay Systems
Quality and User-friendly • Expert Technical Support • Competitive Prices • Expansive Catalogue • Trusted Globally
