EnzyChrom™ Glutamine Assay Kit
Application
- For quantitative determination of glutamine and evaluation of drug effects on glutamine metabolism.
Key Features
- Sensitive and accurate. Use a 20 µL sample. Linear detection range 0.023 – 2 mM glutamine in a 96-well plate assay.
- Convenient. The procedure involves adding a single working reagent, incubation for 40 min at room temperature, adding a stop reagent, and reading the optical density. No 37°C heater is needed.
- High-throughput. Can be readily automated as a high-throughput 96-well plate assay for thousands of samples per day.
Method
- OD565nm
Samples
- Serum, plasma, urine, cell, tissue, etc
Species
- All
Procedure
- 40 min
Size
- 100 tests
Detection Limit
- 23 µM
Shelf Life
- 6 months
More Details
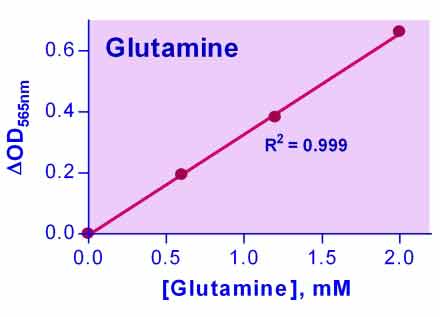
Glutamine is an amino acid synthesized in the muscle that plays major roles in protein synthesis, acid-base balance, and anabolic processes and is utilized for cellular energy and as a carbon source. It is used in the treatment of injury, trauma, and burns, and also as a supplement for muscle growth and post-surgery healing. Simple, direct, and automation-ready procedures for measuring glutamine concentration are very desirable. BioAssay Systems EnzyChrom™ glutamine assay kit is based on the hydrolysis of glutamine to glutamate and colorimetric determination of the product. The intensity of the product color, measured at 565 nm, is proportional to the glutamine concentration in the sample.Does ascorbic acid interfere with the glutamine assay?
Up to 125 mM ascorbic acid in the sample does not interfere significantly with the assay. Samples with higher ascorbic acid concentrations should be diluted to a final ascorbic acid concentration of 125 mM or less. Normal human serum contains around 20-60 mM ascorbic acid, which does not interfere with the assay.
Can I deproteinate my samples with acid?
The assay is based on an enzyme based reaction and acid based deproteination procedures can often interfere with the assay. We do not recommend acid deproteination on samples with this assay.
Ishak Gabra, M. B., et al. (2020). Dietary glutamine supplementation suppresses epigenetically-activated oncogenic pathways to inhibit melanoma tumour growth. Nature Communications. 11(1): 3326. Assay: Glutamine in mouse tissue.
Velickovic, K., et al. (2020). Targeting glutamine synthesis inhibits stem cell adipogenesis in vitro. Cellular Physiology and Biochemistry, 54(5), 917-927. Assay: Glutamine in mouse cell extract.
Qu, Y., et al.(2021). Soybean CHX-type ion transport protein GmSALT3 confers leaf Na + exclusion via a root derived mechanism, and Cl – exclusion via a shoot derived process. Plant, Cell & Environment. 44(3): 856-869. Assay: Glutamine in soybean phloem sap.
Stern, R. A., et al. (2019). Ammonia induces a myostatin-mediated atrophy in mammalian myotubes, but induces hypertrophy in avian myotubes. Frontiers in Sustainable Food Systems. 3: 115. Assay: Glutamine in chicken myotube cultures.
Luo, Z., et al. (2020). Co-delivery of 2-Deoxyglucose and a glutamine metabolism inhibitor V9302 via a prodrug micellar formulation for synergistic targeting of metabolism in cancer. Acta Biomaterialia. 105: 239-252. Assay: Glutamine in mouse and human cancer cell extract.
Tran, T. Q., et al. (2020). α-Ketoglutarate attenuates Wnt signaling and drives differentiation in colorectal cancer. Nature Cancer. 1(3): 345-358. Assay: Glutamine in human colonic tissue.
Dill, Veronika, et al (2019). Investigation of cell culture conditions for optimal foot-and-mouth disease virus production. BMC Biotechnology 19.1: 33. Assay: Glutamine in hamster cells.
Yang, Ying, et al (2019). MiR-135 suppresses glycolysis and promotes pancreatic cancer cell adaptation to metabolic stress by targeting phosphofructokinase-1. Nature communications 10.1: 809. Assay: Glutamine in mice tissues.
Bar-Sagi, Dafna, Cosimo Commisso, and Rengin G. Soydaner-Azeloglu (2018). Cancer diagnostics, therapeutics, and drug discovery associated with macropinocytosis. U.S. Patent No. 9,983,194. Assay: Glutamine in mouse cells.
Basit, Farhan, et al (2018). human dendritic cell subsets undergo distinct metabolic reprogramming for immune response. Frontiers in immunology 9:2489. Assay: Glutamine in human cells.
Ma, Huanrong, et al (2018). Inhibition of SLC1A 5 sensitizes colorectal cancer to cetuximab. International journal of cancer 142.12: 2578-2588. Assay: Glutamine in human cells.
Shen, Guangsi, et al (2018). GOLM1 Stimulation of Glutamine Metabolism Promotes Osteoporosis via Inhibiting Osteogenic Differentiation of BMSCs. Cellular Physiology and Biochemistry 50.5: 1916-1928. Assay: Glutamine in mice cells.
Stern, R. A. (2018). Myogenic Response to Ammonia Differs Between Avian and Mammalian Species. Assay: Glutamine in mouse & chicken cells.
Wang, Yun-Qian, et al (2018). Sirtuin5 contributes to colorectal carcinogenesis by enhancing glutaminolysis in a deglutarylation-dependent manner. Nature communications 9.1: 545. Assay: Glutamine in human cells.
Pugh, Jamie N., et al (2017). Glutamine supplementation reduces markers of intestinal permeability during running in the heat in a dose-dependent manner. European journal of applied physiology 117.12: 2569-2577. Assay: Glutamine in human plasma.
Chen, Jianmin, et al (2016). The impact of glutamine supplementation on the symptoms of ataxia-telangiectasia: a preclinical assessment. Molecular neurodegeneration 11.1: 60. Assay: Glutamine in mice blood.
Kempaiah, Prakasha, et al (2016). Reduced Hsp70 and Glutamine in Pediatric Severe Malaria Anemia: Role of hemozoin in Suppressing Hsp70 and NF-kB Activation. Molecular Medicine 22.1: 570-584. Assay: Glutamine in human plasma.
Zhul, M et al (2015). The effects of acute oral glutamine supplementation on exercise-induced gastrointestinal permeability and heat shock protein expression in peripheral blood mononuclear cells. Cell Stress and Chaperones 20(1): 85-93. Assay: Glutamine in human plasma.
Behari, J et al (2014) b-Catenin Links Hepatic Metabolic Zonation with Lipid Metabolism and Diet-Induced Obesity in Mice. The American Journal of Pathology. 184:12. Assay: Glutamine in mice liver cells.
Behousiar, A et al (2014). FIBS-enabled noninvasive metabolic profiling. Journal of Visualized Experiments (84):e51200. Assay: Glutamine in hamster cell lines.
Zhang. M et al (2014), Oral cancer cells may rewire alternative metabolic pathways to survive from siRNA silencing of metabolic enzymes. BMC Cancer 14:223. Assay: Glutamine in human oral keratinocytes.
Zhul, MN et al (2014). Effects of oral glutamine supplementation on exercise-induced gastrointestinal permeability and tight junction protein expression. Journal of Applied Physiology. 116(2):183-191. Assay: Glutamine in human intestinal tissue.
Behjousiar A et al (2012). In situ monitoring of intracellular glucose and glutamine in CHO cell culture. PLoS One 7(4):e34512. Assay: Glutamine in hamster CHO cells.
Lin TC et al (2012). Autophagy: Resetting glutamine-dependent metabolism and oxygen consumption. Autophagy 8(10):1477-93. Assay: Glutamine in mouse mouse embryonic fibroblasts.
Qi S et al (2012). Comparison of the metabolic profiling of hepatitis B virus-infected cirrhosis and alcoholic cirrhosis patients by using (1) H NMR-based metabonomics. Hepatol Res 42(7):677-85. Assay: Glutamine in human serum.
Tang N et al (2012). Stable overexpression of arginase I and ornithine transcarbamylase in HepG2 cells improves its ammonia detoxification. J Cell Biochem. 113(2):518-27. Assay: Glutamine in human hepatocellular carcinoma cells.
Liang, Y et al (2011). Additive effect of tetramethylpyrazine and deferoxamine in the treatment of spinal cord injury caused by aortic cross-clamping in rats. Spinal Cord 49(2):302-6. Assay: Glutamate in rat tissue.
Rink, C et al (2011). Oxygen-inducible glutamate oxaloacetate transaminase as protective switch transforming neurotoxic glutamate to metabolic fuel during acute ischemic stroke. Antioxid Redox Signal 14(10):1777-85. Assay: Glutamate in rat Brain tissue.
Yang LM, Blount P (2011). Manipulating the permeation of charged compounds through the MscL nanovalve. FASEB J 25(1):428-34. Assay: Glutamate in Bacteria E. coli glutamate efflux.
To find more recent publications, please click here.
If you or your labs do not have the equipment or scientists necessary to run this assay, BioAssay Systems can perform the service for you.
– Fast turnaround
– Quality data
– Low cost
Please email or call 1-510-782-9988 x 2 to discuss your projects.
$549.00
For bulk quote or custom reagents, please email or call 1-510-782-9988 x 1.
Orders are shipped the same day if placed by 2pm PST
Shipping: On Ice
Carrier: Fedex
Delivery: 1-2 days (US), 3-6 days (Intl)
Storage: -20°C upon receipt
Related Products
You may also like…
| Name | SKU | Price | Buy |
|---|---|---|---|
| EnzyChrom™ Glutamate Assay Kit | EGLT-100 | $489.00 |
Why BioAssay Systems
Quality and User-friendly • Expert Technical Support • Competitive Prices • Expansive Catalogue • Trusted Globally
